3-Color Crystal Digital PCR™ assays for HPV16 and HPV18 detection
Sensitive and specific detection of HPV16 and HPV18 in human DNA
Human papillomavirus (HPV), a sexually transmitted infection, leads to more than 5% of cancers in the world [1]. More than a dozen HPV subtypes are regarded as high-risk due to their direct role in the transformation of cells and an increased probability of cervical and oropharyngeal cancers. Among these subtypes, the HPV16 and HPV18 are the most prevalent, detected in approximately 70% of cervical cancers and in 90% of oropharyngeal cancers [1]. HPV16 and 18 immortalize and transform cells by integrating stretches of their viral sequences (including the oncogenic E6 and E7 regions) into the cellular genome. Early HPV detection using sensitive and specific molecular techniques capable of distinguishing high-risk HPV types from low-risk HPV types is essential to the treatment of precancerous lesions and the prevention of cervical cancer progression. Here, we show how a 3-color Crystal Digital PCRTM assay (Table 1) on the NaicaTM System reliably detects and quantifies HPV16, HPV18 and the human reference gene ALB within a human DNA sample (Figure 1).

Using an optimized 3-color Crystal Digital PCR HPV assay, no false positives were detected in 36 replicates containing human wild-type genomic DNA only (final concentration 400 cp/µl), resulting in a limit of blank of zero. Serial dilutions using synthetic HPV16 and HPV18 DNA sequences in a background of human wild-type DNA (final concentration 400 cp/µl) were assessed in triplicate. The targeted HPV16 and HPV18 sequences were detected with a 95% confidence level at a final concentration down to 0.2 copies per microliter, representing a mutant allele frequency of 0.05% (Figure 2).
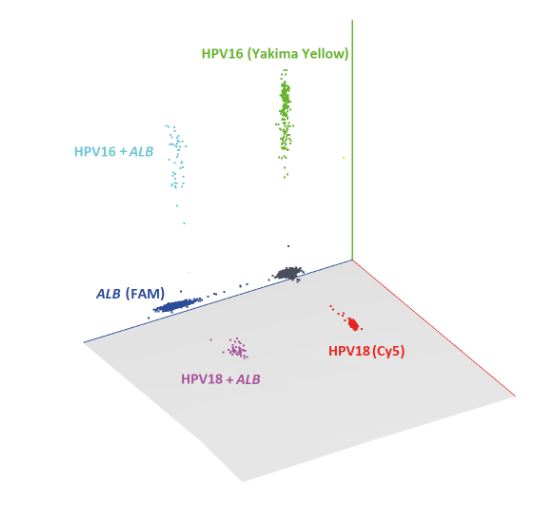
Figure 1: 3D dot plot generated by Crystal Miner software for simultaneous display of HPV16, HPV18 and ALB reference gene detection. Fluorophore names are indicated in parentheses.
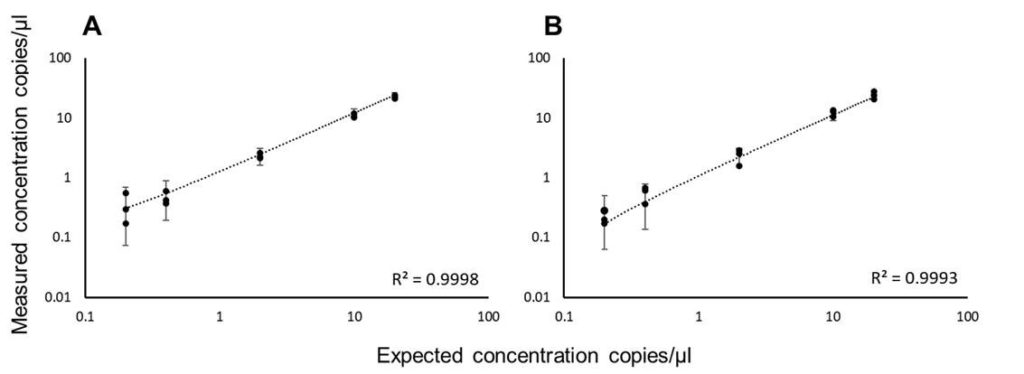
Figure 2: Linearity and sensitivity of the 3-color Crystal Digital PCR assay for the detection of (A) HPV16 and (B) HPV18 targets. Serial dilutions ranging from 20 to 0.2 cp/µl (500 to 5 copies per 25µl reaction) of HPV16 and HPV18 in a background of 400 cp/μl of WT DNA (10,000 copies per 25µl reaction) were assayed in triplicates. The vertical bars represent the theoretical 95% confidence intervals around the experimental mean values. A total of 0.2 cp/µl of HPV16 and HPV18 (5 copies per 25µl reaction) was reliably detected.
Robust HPV16 and HPV18 detection in patient samples
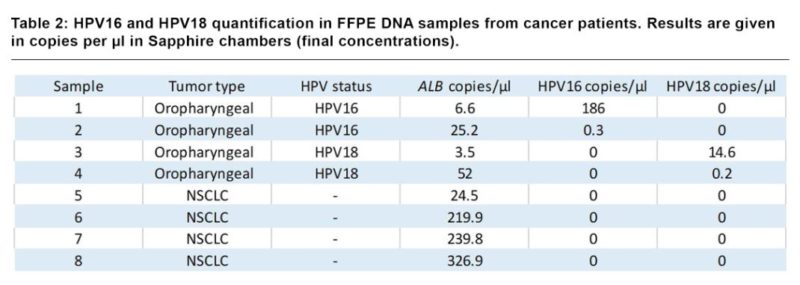
DNA samples extracted from four formalin-fixed paraffin-embedded (FFPE) tumor tissues of oropharyngeal cancer patients and four FFPE of non-HPV infected non-small cell lung cancer (NSCLC) patients were analyzed using the 3-color Crystal Digital PCR HPV assay (Table 2). Oropharyngeal FFPE samples were previously characterized using real-time PCR [2]. All four HPV16 and HPV18 positive samples were detected using the 3-color Crystal Digital PCR HPV assay whereas non-HPV samples were found negative for HPV.
[1]: T. A. Berman et J. T. Schiller, “Human papillomavirus in cervical cancer and oropharyngeal cancer: One cause, two diseases”, Cancer, vol. 123, no 12, p. 2219‑2229, 2017. [2]: Melkane AE, Mirghani H, Aupérin A, Saulnier P, Lacroix L, Vielh P, CasiraghiO, Griscelli F, Temam S. HPV-related oropharyngeal squamous cell carcinomas: a comparison between three diagnostic approaches. Am J Otolaryngol. 2014 Jan-Feb;35(1):25-32.
FFPE samples were the kind gift of Ludovic Lacroix, Institut Gustave Roussy.
Application Note Highlights
- The naica® 3-color Crystal Digital PCR system enables the simultaneous detection and quantification of HPV16 and HPV18, as well as a human reference gene.
- HPV16 and HPV18 are reliably detected and quantified at 0.05% mutant allele frequencies in a background of wild-type DNA of 10,000 copies per 25µl reaction with a 95% confidence level while no false positives were detected.
- In serial dilution experiments, the 3-color HPV assay displays an excellent correlation between the expected and measured HPV16 and HPV18 concentrations.
- 3-color Crystal Digital PCR successfully detects HPV16 and HPV18 in DNA extracted from oropharyngeal cancer patient FFPE tumor samples.
To learn more about digital PCR, please visit Stilla Technologies’ Learning Center.